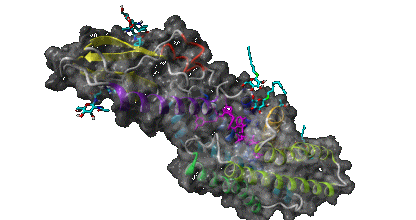
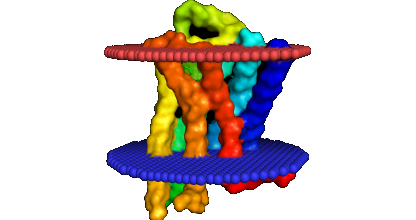
Results Page
Status:
Task completed successfully.
Submission info:
| Task mode - Auto |
| Username: | pledor | Set of templates: | inactive |
| Task token: | f629affe700c4507b4d2934e9eb77ff9 | Lysozyme included: | no |
| Date of submission: | 11/15/12 09:53 AM | Run time: | 0d 17h 55m |
| Date started: | 11/15/12 09:53 AM | Branch cutoff: | 0.01 |
| Date finished: | 11/16/12 03:49 AM | Min e-value: | 1e-100 |
| Query length: | Unknown | Max e-value: | 1e-10 |
|
| Query sequence: |
| File missing: targ_sequence. Sorry. |
Results:
| Template selection: |
| File missing: target_scores.csv. Sorry. |
| Alignments generated with selected templates: |
| File missing: alignments_scores.csv. Sorry. |
| Final Alignment: |
| File missing: alignment_convergence.log. Sorry. |
| Final Alignment in the PIR format: |
| File missing: align.ali. Sorry. |
| Loops to be refined (Modeller): |
| File missing: modeller-loops.csv. Sorry. |
| Generated Modeller Models: |
| File missing: modeller/modeller-models.csv. Sorry. |
| Loops to be refined (Rosetta): |
| File missing: rosetta/rosetta-loops.csv. Sorry. |
| Generated Rosetta Models (sorted by BCL score): |
| File missing: rosetta/rosetta-models_bcl.csv. Sorry. |
| Generated Rosetta Models (sorted by RosettaMP score): |
| File missing: rosetta/rosetta-models_mp.csv. Sorry. |
| Generated Rosetta Models (sorted by Rosetta total energy): |
| File missing: rosetta/rosetta-models.csv. Sorry. |
| Sequences in the query profile. Sequences are sorted by e-value in descending order. The first sequence in the file is your query sequence. |
| File missing: targ_family_selected.fasta. Sorry. |
| Anchored realignment - a log file. |
| File missing: check_alignment.log. Sorry. |
| Modeller run script (1) |
| File missing: model.py. Sorry. |
| Modeller run script (2) |
| File missing: myloop.py. Sorry. |
| Modeller Run - a log file. |
| File missing: modeller/modeller.log. Sorry. |
| Rosetta Run - a log file. |
| File missing: rosetta/rosetta.log. Sorry. |
| Alignments & Models - zip file. |
| File missing: results.zip. Sorry. |
Thank you for using GPCRM Structure Modelling Server!
Any questions? Contact us: support|biomodellab.eu